Mapping the brain
26 Aug 2011
The brain of a mouse measures only 1 cubic centimetre in volume. But when neuroscientists at Harvard's Center for Brain Science slice it thinly and take high-resolution micrographs of each slice, that tiny brain turns into an exabyte of image data. That's 1018 bytes, equivalent to more than a billion CDs.
What can you do with such a gigantic, unwieldy data set? That's the latest challenge for Hanspeter Pfister, the Gordon McKay Professor of the Practice of Computer Science at Harvard's School of Engineering and Applied Sciences (SEAS).
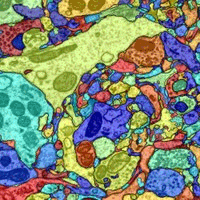 |
| Pfister and his colleagues have developed an algorithm that automatically detects the boundaries of cells in a sheet of tissue, allowing cells to be matched up across brain slices. Image courtesy of Amelio Vasquez-Reina. |
Pfister, an expert in high-performance computing and visualisation, is part of an interdisciplinary team collaborating on the Connectome Project at the Center for Brain Science. The ambitious Connectome Project aims to create a wiring diagram of all the neurons in the brain, and neuroscientists have developed innovative techniques for automatically imaging slices of mouse brain, yielding terabytes of data so far.
Pfister's system for displaying and processing these images would be familiar to anyone who has used the Google Maps service. Because only a subsection of a very large image can be displayed on a screen, only that viewable subsection is loaded. Drag the image around, zoom in or out, and more of the image is displayed on the fly.
This ''demand-driven distributed computation'' is the central idea behind Pfister's work, which recently won him a Google Faculty Research Award.
With new techniques producing ever-larger data sets in science, the problem is hardly unique to the Connectome Project. Pfister is certain that the tools his research group develops can be used by scientists working with large image data sets in any field - for example, astronomers processing radio telescope images or earth scientists analysing atmospheric data.
Even for the Connectome, the ultimate tool will be much more than ''Google Maps for the brain.'' The project has two crucial new features, says Pfister. First, the mouse brain must be reconstructed in three dimensions, so one can quickly flip through the stack of images in addition to zooming and panning across it. The images, each representing a 30-nanometre-thick brain slice, can also be stacked together and viewed from the side, along the same axis they were sliced.
Additionally, the system developed by Pfister's research team extends the demand-driven principle to image analysis and processing. If a neuroscientist wants to automatically align images, adjust the brightness of an image, or run an automatic cell boundary detector over a portion of the images, for example, these operations are also computed on the fly, so there is no need to wait for the entire stack of images to be processed before the relevant subsection is viewable. Pfister compares it to ''a very specialised version of Adobe Photoshop software.''
Indeed, the existing tools are capable of far more advanced image processing. Since the collaboration began in 2007, computer scientists in Pfister's group have created algorithms to automatically detect cell boundaries, a process known as segmentation, and to connect related cells in three dimensions, a process known as reconstruction. Previously, neuroscientists did this by hand.


















