Lollipops and ice fishing: Molecular rulers used to probe nanopores
04 May 2010
Using a pair of exotic techniques, including a molecular-scale version of ice fishing, a team of researchers working at the National Institute of Standards and Technology (NIST) have developed methods to measure accurately the length of "nanopores," the miniscule channels found in cell membranes. The "molecular rulers" they describe in a recent paper* could serve as a way to calibrate tailor-made nanopores-whose diameters on average are nearly 10,000 times smaller than that of a human hair-for a variety of applications such as rapid DNA analysis.
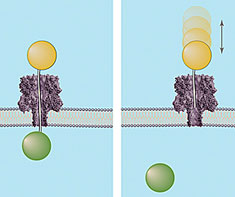 |
| Graphic depicting how the "ice fishing" method determines the distance across a membrane nanopore. Both images show DNA strands of known lengths topped by a polymer cap (orange sphere) being driven through the nanopore. If the DNA strand is long enough to completely transverse the channel (left), it will "hook" a circulating polymer (green sphere) on the other side of the membrane and define the nanopore's length. If not long enough, the DNA probe will bounce out of the pore (right). |
Graphic depicting how the "ice fishing" method determines the distance across a membrane nanopore. Both images show DNA strands of known lengths topped by a polymer cap (orange sphere) being driven through the nanopore. If the DNA strand is long enough to completely transverse the channel (left), it will "hook" a circulating polymer (green sphere) on the other side of the membrane and define the nanopore's length. If not long enough, the DNA probe will bounce out of the pore (right).
Studies at NIST and other research institutions have shown that a single nanometer-scale pore in a thin membrane can be used as a "miniature analysis laboratory" to detect and characterize individual biological molecules such as DNA or toxins as they pass through or block the passage. Such a system could potentially fit on a single microchip device, for a wide variety of applications. However, making the mini-lab practical requires an accurate definition of the dimensions and structural features of the nanopore.
In new experiments, researchers from NIST and the University of Maryland first built a membrane-a bilayer sheet of lipid molecules-similar to that found in animal cells.
They "drilled" a pore in it with a protein** designed specifically to penetrate cell membranes. When voltage is applied across the membrane wall, charged molecules such as single-stranded DNA are forced into the nanopore. As the molecule passes into the channel, the ionic current flow is reduced for a time that is proportional to the size of the chain, allowing its length to be easily derived.
If a chain is long enough to reach the narrowest part of the nanopore-known as the pinch point-the force of the electrical field behind it will push the molecule on through the rest of the channel. Exploiting this characteristic, the NIST/Maryland team developed a DNA probe method to measure the distances from the openings on each side of the membrane to the pinch point, and in turn, the entire length of the nanopore by adding the two measurements together.

















